Table of Contents
3. Tentative correspondences and RANSAC
Preliminaries
Before we start, we'll need a function pts=detect_and_describe(img,detpar,descpar) connecting the previously implemented functions together, such that it detects, geometrically and photometrically normalizes and describes feature points in the input image img. Detector parameters will be passed in structure detpar:
- for Hessian detector::
detpar.type='hessian'; detpar.sigma % sigma for derivatives detpar.threshold
- for Harris detector:
detpar.type='harris'; detpar.sigmad % sigma for derivatives detpar.sigmai % sigma for integration detpar.threshold
- for Hessian detector in scale-space
detpar.type='sshessian'; detpar.threshold
- for MSER detector
detpar.type='mser'; detpar.min_margin % requested stability detpar.min_size % area of the smallest region (in pixels) detpar.max_area % area of the biggest region (in relation to image area, in range <0, 1>)
Parameters for normalization and description will be passed in structure descpar
descpar.type % type of the description ('dct' nebo 'ghisto' nebo 'sift') descpar.ps % size of patch descpar.ext % size of feature point neighborhood in canonical coordinate systemextra for DCT descriptor:
descpar.num_coeffs % number of coefficient in zig-zag orderextra for ghisto:
descpar.num_bins % number of bins in one ax of histogram
The structure pts on the output is in the format of function affnorm (array x,y,a11,a12,a21,a22,patch) and photonorm (mean,std) from last labs, for each point <latex>i</latex> adds array pts(i).desc with the description as a column vector. You can find function detect_and_describe here.
Tentative correspondences
After detection and description, which run on each image of the scene separately, we need to find correspondences between the images. The first phase is looking for the tentative correspondences (sometimes called “matches”). A tentative correspondence is a pair of features with descriptions similar in some metric. From now on, we will refer to our images of the scene as the left one and the right one. The easiest way is to establish tentative correspondences is to find all distances between the descriptions of features detected in the left and the right image. We call this table a “distance matrix”. We will compute the distance in Euclidean space. Let's try several methods for correspondence selection:
- Tentative correspondences of mutually nearest descriptions are created, when the nearest description in the left image for description A from right image is description B and at the same time, description A is the closest description in the right image to description B from the left image.
- For stable pairing in each step the algorithm finds globally closest descriptions A and B. It creates tentative correspondence and descriptions A and B are excluded from finding in next steps (for instance with setting infinity value to right place in distance matrix). The iterations continues while there are any active points left (values smaller than infinity).
- A different procedure is to find the two closest descriptions in the right image for each description from the left image and then test, if the ratio of first/second closest description is smaller than a threshold (use e.g. 0.8). If so, the descriptions correspond and it is guaranteed, that there are no confusing descriptions which would be too close in the description space. This method can be used for well discriminative description - for instance SIFT.
The output of finding tentative correspondences are pairs of indices of features from the left and the right image with the closest descriptions.
- write a function
corrs=match(pts1, pts2, par), which finds correspondences between pointspts1andpts2with method specified in theparstructure. Description of each point $i$ is saved in arraypts1(i).descresp.pts2(i).desc. The method is specified in the fieldpar.methodas a string. For each method, ignore tentative correspondences with distance greather than thresholdpar.threshold.
par.method = 'mutual'; % for mutual nearest method 'stable'; % for stable pairing 'sclosest'; % for "first/second closest" method par.threshold % threshold, pairs with distance greater than this will be ignoredOutput will be a 2xN matrix containing the indices of tentative correspondences, where the first row corresponds to indices in
pts1 and second row to indices in pts2.
RANSAC and the geometric model of the scene
The tentative correspondences from the previous step usually contain many non-corresponding points, so called outliers, i.e. tentative correspondences with similar descriptions that do not correspond to the same physical object in the scene. The last phase of finding correspondences aims to get rid of the outliers. We will assume images of the same scene and search a model of the undergoing transformation. The result will be a model of the transformation and a set of the so called inliers, tentative correspondences that are consistent with the model.
The model of a planar scene
The relation between the planes in the scene is discussed in the course Digital Image. In short:
The transformation between two images of a plane observed from two views (left and right image) using the perspective camera model is a linear transformation (called homography) of 3-dimensional homogeneous vectors (of corresponding points in the homogenoeous coordinates) represented by a regular $3\times 3$ matrix $\mathbf{H}$:
$$ \lambda\left[\!\begin{array}{c}x'\\y'\\1\end{array}\!\right]= \left[\begin{array}{ccc}h_{11}&h_{12}&h_{13}\\h_{21}&h_{22}&h_{23}\\h_{31}&h_{32}&h_{33}\end{array}\right] \left[\!\begin{array}{c}x\\y\\1\end{array}\!\right]% $$
For each pair of corresponding points $(x_i,y_i)^{\top} \leftrightarrow (x'_i,y'_i)^{\top}$ it holds:
$$\begin{array}{lcr} x' (h_{31}\,x + h_{32}\,y + h_{33}) - (h_{11}\,x + h_{12}\,y + h_{13}) & = & 0 \\ y' (h_{31}\,x + h_{32}\,y + h_{33}) - (h_{21}\,x + h_{22}\,y + h_{23}) & = & 0 \end{array}$$
Let us build a system of linear constraints for each pair:
$$ \underbrace{\left[\begin{array}{r@{\ }r@{\ }rr@{\ }r@{\ }rr@{\ \ }r@{\ \ }r} -x_1&-y_1&-1&0&0&0&x_1'x_1&x'_1y_1&x_1'\\0&0&0&-x_1&-y_1&-1&y_1'x_1&y'_1y_1&y_1'\\ -x_2&-y_2&-1&0&0&0&x_2'x_2&x'_2y_2&x_2'\\0&0&0&-x_2&-y_2&-1&y_2'x_2&y'_2y_2&y_2'\\ &\vdots&&&\vdots&&&\vdots&\\ -x_n&-y_n&-1&0&0&0&x_n'x_n&x'_ny_n&x_n'\\0&0&0&-x_n&-y_n&-1&y_n'x_n&y'_ny_n&y_n'\\ \end{array}\right]}_\mathbf{C} \left[\begin{array}{c}h_{11}\\h_{12}\\h_{13}\\h_{21}\\h_{22}\\h_{23}\\h_{31}\\h_{32}\\h_{33}\end{array}\right]=0 $$
This homogeneous system of equations has 9 degrees of freedom and we get one non-trivial solution as a right nullspace of the matrix <latex>\mathbf{C}</latex>. For a unique solution we need at least 8 linearly independent rows of matrix <latex>\mathbf{C}</latex>, i.e. 8 constraints from 4 corresponding points in a general position (no triplet of points can lie on a line!).
- implement a function
H=u2h(u)that gets a matrix of 4 corresponding pairs of points in homogeneous coordinates and computes the homography <latex>\mathbf{H}</latex>. In case that some of the triplets will lie close to a line, it returns an empty matrix []. The input u is a 6×4 matrix as shown below, where xi,yi are point coordinates in the left image and x'i,y'i in the right image. :
$$ u = \left[\begin{array}{cccc} x_1 & x_2 & x_3 & x_4 \\ y_1 & y_2 & y_3 & y_4 \\ 1 & 1 & 1 & 1 \\ x_1' & x_2' & x_3' & x_4' \\ y_1' & y_2' & y_3' & y_4' \\ 1 & 1 & 1 & 1 \end{array}\right] $$
- write a function
dist=hdist(H,u), which gets a 6xN matrix u of N pairs of points in homogeneous coordinates and returns a vector dist (1xN) of squared distances of the points <latex>\mathbf{H} (x_i, y_i,1)^\top</latex> and <latex>(x'_i, y'_i,1)^\top</latex> in Euclidean space.
RANSAC
To find a correct model of the scene – a projective transformation between two planes, we need at least 4 inlier point pairs (not in a line). One way of picking such points in a robust way from all the pairs of acquired tentative correspondences is the RANSAC algorithm of M.Fischler and R.C.Bolles.
In our task, the RANSAC algorithm will have this form:
- Uniformly sample 4 corresponding pairs. You can use the provided function
sample.m. - Compute the homography $\mathbf{H}$ out of the sampled point pairs.
- Compute the support of the found model. If the transformation $\mathbf{H}$ was computed from an all-inlier sample, all inlier points from the left image $(x_i,y_i)$ shall be projected by $\mathbf{H}$ to the corresponding points $(x'_i,y'_i)$ in the other image. The support of the model is quantified by the number of points consistent with the model, i.e. the number of correspondences with distance (function
hdist) of transformed point from the left image and the corresponding point in the right image smaller than a predefined threshold (e.g. 1-3 pixels). - If the support of the model is the biggest so far, we keep this so-far-the-best homography $\mathbf{H}^*$ and the corresponding set of inliers, i.e points that supported this model.
- We iterate points 1-4 and keep track of so-far-the-best $\mathbf{H}^*$ transformation.
Set the number of iterations to 1000 in the beginning and debug the RANSAC algorithm on a simple pair of images. A more sophisticated stopping criterion is used in practice. It is based on the probability estimate of finding an all-inlier sample (of size 4 in our case) under the assumptions of iteratively estimated fraction of inliers. The stopping criterium is implemented in functionnsamples.m. It computes total number of iterations required for a given confidence conf:
num_samples = nsamples(number_of_inliers, number_tentative_correspondences, sample_size, conf);where number_of_inliers is iteratively estimated at each observation of a new so-far-the-best model, i.e. for each new so-far-the-best model we recompute the number of steps required by
nsamples.
- use the provided functions to implement s function
[Hbest,inl]=ransac_h(u,threshold,confidence)that robustly estimates the homography $\mathbf{H}^*$ and a set of inliers inl. Input u is a 6xN matrix of corresponding points in homogeneous coordinates (similar to functionhdist). The parameter threshold is the maximum distance of reprojected points (computed viahdist) that will be marked as inliers (pairs consistent with the model $\mathbf{H}^*$). The parameter confidence is a value from the range (0,1), the requested confidence of getting a correct sample (a good value for testing is for example 0.95).
The output is the best model $\mathbf{H}^*$ and an array inl (1xN), where 0 represents an outlier and 1 represents an inlier.
Position of two perspective cameras, epipolar geometry
To verify the tentative correspondences for other than planar scenes, we will need more general relation between two points from two cameras. The geometric relation of a pair of uncalibrated cameras is called the epipolar geometry. The epipolar geometry is discussed in the course of Geometry of Computer Vision and Graphics, for an initial information you can read the corresponding wikipedia page or listen to a dedicated song :). For our problem, it is important that the epipolar geometry is a geometric relation of corresponding points $x_L$ a $x_R$, images of a physical point $X$ in the scene observed by a pair of perspective cameras:
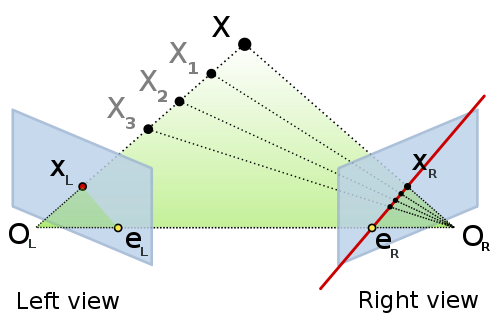 |
| Epipolar geometry, source: wikimedia illustration by Arne Nordmann |
This relation can be written as:
$$ x^\top_L \mathbf{F}\, x_R = 0 $$
The epipolar geometry is represented by a matrix $\mathbf{F}$, called the fundamental matrix. To find a fundamental matrix we need at least 7 corresponding points. Our RANSAC implementation from the previous section has to be modified in two ways: We'll need a function u2f7.m that will replace the estimation of homography by estimation of the fundamental matrix $\mathbf{F}$ and a function fds.m that will replace the function hdist that estimates the distance of the reprojected points, here the distance of the corresponding point from an epipolar line (image of the ray through the point in the other camera). These functions have the same parameters as u2h and hdist, so you can just replace matrix $\mathbf{H}$ by matrix $\mathbf{F}$. Function u2f7 returns from one (result is a 3×3 matrix) up to three solutions (result is a 3x3xnumber_of_solutions matrix), thus it is required to modify the verification of the model to pick the best of returned solutions.
- use the provided functions to implement a function
[Fbest,inl]=ransac_f(u,threshold,confidence)that estimates the fundamental matrix $\mathbf{F}^*$ and a set of inliers inl. Input u is a 6xN matrix of corresponding points in homogeneous coordinates. Parameter threshold defines the maximal distance of points for functionfdsto be labeled as inliers, consistent with the epipolar geometry given by $\mathbf{F}^*$. The parameter confidence is value from the range (0,1), requested probability of getting a correct sample (for testing use the value 0.95).
The result will be the best model found $\mathbf{F}^*$ and an arrayinl(1xN), where 0 represents an outlier and 1 represents an inlier.
What should you submit?
Implement functions match.m, u2h.m, hdist.m, ransac_h.m, ransac_f.m and submit them in task 04_corr together with all non-standard functions you have created.
Choose a combination of detector/description for each object in your dataset. Run the detectors and generate descriptions for all images using function detect_and_describe. Save the results (structure pts) into file obj%d_view%d.mat together with your configuration of the detector (detpar), normalization and description (descpar). Objects and views numerate from 0 (e.g. for object 1 image 3 save the results: structures pts, detpar and descpar into file obj1_view3.mat, for object 0 image 0 in file obj0_view0.mat, etc.).
Find tentative correspondences for each object and all pairs of views to the object. The result will be a matrix TC of cells 5×5, where cells will contain arrays corrs form function match. We will assume, that the process is symmetric (2→1 is same as 1→2) and generate only upper triangle (index of first image as row and second image as column) without diagonal. Leave other cells empty. Save matrix TC together with configuration par of the method match into file obj%d_tc.mat.
At the end run both RANSACs on generated tentative correspondences for each object and save the results into matrices H, F, inlH and inlF of cells 5×5. Save the cell matrices H, F, inlH and inlF into file obj%d_results.mat together with parameters thresh_h, thresh_f, conf_h, conf_f of function ransac_h and ransac_f.
Pack the all results into an archive together with your images from task 02_data named (username)_obj(0-2)_view(0-4).jpg; and submit into task 04_results. To save some space you can remove the field patch from structure pts using function rmfield.
Testing
To test your code, download corr_test.zip and unpack it into directory in MATLAB paths (or put it into directory with your code) and execute. Compare your results with ours.